This web page was produced as an assignment for Gen677 at UW-Madison Spring 2013
Conclusions
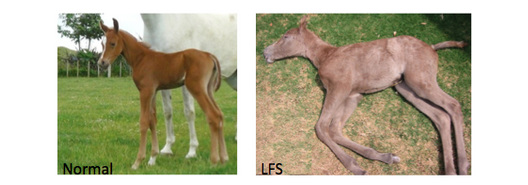
This page will outline the experiments and future directions for research into MYO5a function that I have designed after completing my research into Lavender Foal Syndrome. There are two areas of particular interest that I came across in my research. The first is the uncharacterized low complexity regions found in the coiled coil domain of MYO5a. These highly conserved regions have no known function but could potentially be an uncharacterized protein domain or motif. The second is the fact that MYO5a function is poorly understood in neurons [1].Neurological defects in animals with MYO5a cause death but the actual mechanism resulting in the neurologic symptoms are not well characterized.
Experiments
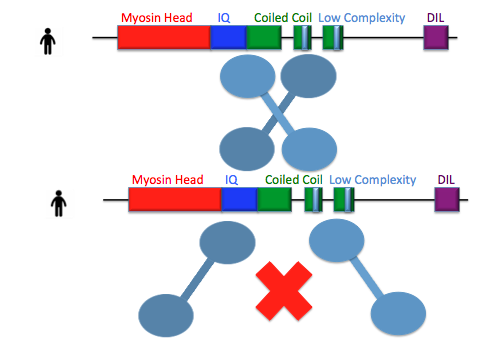
The first areas of MYO5a that I want to investigate are the low complexity regions. These regions are very well conserved in all vertebrates analyzed suggesting that they have some function important for the protein. These regions are found in the coiled coil domain of MYO5a which is responsible for protein dimerization [1], leading me to my first hypothesis:
Hypothesis 1: The low complexity region in MYO5a is important for dimerization.
|
Experiment 1: The first step towards answering this questions it to see if the amino acid sequence of these low complexity regions are conserved. I achieved this by performing a Clustal Omega alignment of the sites [4][5]. This alignment showed that the low complexity regions have 100% amino acid conservation in vertebrates. The next step I propose is a mutagenesis of an amino acid residue in the low complexity region, for example of Leucine resides found at location 1380. If the low complexity region is in fact important for dimerization then the mutagenesis should prevent dimerization. Failure to dimerize could be assayed using TAP-tag to see if the mutated protein is able to still interact with MYO5a.
|
The second area that I want to explore in MYO5a is the possibility of novel proteins that interact with MYO5a but have not yet been characterized. The function of MYO5a in neurons in of critical importance but is poorly characterized [1]. MYO5a function in melanocytes is believed to be similar and is much simpler to study, possibly providing clues to MYO5a functions and interactions in neurons. One of the key pieces of information about MYO5a that is not well understood is the network of proteins it interacts with in the neurons [1]. When thinking about this I began to wonder if there were proteins as yet unidentified that interact with MYO5a in melanocytes. This lead me to my second hypothesis:
Hypothesis 2: There are unidentified proteins important for pigmentation that bind to Myo5a.
Hypothesis 2: There are unidentified proteins important for pigmentation that bind to Myo5a.
|
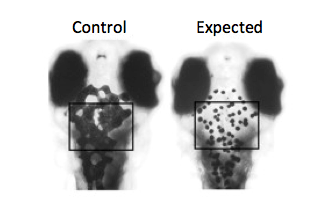
Experiment 2: I propose to study this hypothesis in zebrafish melanocytes. Zebrafish are transparent early in development making any melanocyte phenotype easy to assay. I would first do a genome wide screen using high-throughput RNAi to identify any proteins involved in pigmentation. I would classify this as any protein that produces a melanocyte unable to distribute its melanosomes out along its dendrites, leaving them clustered at the center of the cell, as a result of the RNAi. I would then take the list of proteins that produce the desired melanocyte phenotype and sort them based on gene ontology looking for transport proteins. Finally, I would take any identified transport proteins and test them for MYO5a protein interaction using TAP-tag.
|
Future Directions
If I were to find a novel transport protein interacting with MYO5a in zebrafish the next step would be to knock out the identified protein and MYO5a in zebrafish to see if the melanocyte phenotype was reproducible and the same in both cases. If it was reproduced in both this would indicate that the identified protein does interact with MYO5a in melanocytes.

Figure 4. Left - normal mouse[7]. Right - Mouse with MYO5a mutation[8]
After verifying the phenotype of any identified transport porteins I would want to move the experiments into a mammal model organism. The mouse would be an excellent candidate as MYO5a mutants are already know in this species and a phenotype could be compared between known MYO5a mutations and a knockout of any proteins identified in zebrafish [9]. The hope would be that knocking out an identified novel melanocyte transport protein would result in neurological defects similar to those seen in MYO5a mutants. If this were to happen it would be possible that the identified protein is important to MYO5a function in neurons, moving one step closer to understanding how MYO5a functions in neurons and causes disease. Once MYO5a activity in neurons is completely understood the search can begin for an effective treatment for those suffering from MYO5a mutation neurological diseases such as Lavender Foal Syndrome.
Below is my final presentation outlining my findings and proposed experiments.
| final_presentation_file.pdf | |
| File Size: | 7663 kb |
| File Type: | |
References
[1] Desnos C, Huet S, Darchen F. (2007). "Should I stay or should I go?": myosin V function in organelle trafficking. Biol Cell 99:411-423.
[2] New Foal for 2011-2012 Season. Delight Arab Horse Stud. Retrieved May 14, 2013 from http://delightarabhorsestud.homestead.com/Foals_.html
[3] Lavender Foal Syndrome (LFS). Animal Genetics UK. Retrieved May 14, 2013 from http://animalgenetics.eu/Equine/equine-genetic-disease/lfs.html
[4] Dereeper A., Audic S., Claverie J.M., Blanc G. BLAST-EXPLORER helps you building datasets for phylogenetic analysis. BMC Evol Biol. 2010 Jan 12;10:8. (PubMed)
[5] Dereeper A., Guignon V., Blanc G., Audic S., Buffet S., Chevenet F., Dufayard J.-F., Guindon S., Lefort V., Lescot M., Claverie J.-M., Gascuel O. Phylogeny.fr: robust phylogenetic analysis for the non-specialist Nucleic Acids Research. 2008 Jul 1; 36 (Web Server Issue):W465-9. Epub 2008 Apr 19. (PubMed)
[6] Luo, N. et al. (2012) Evidence for a role of inositol polyphosphate 5-phosphatase INPP5E in cilia formation in zebrafish. Vision Research. 75:98-107.
[7] DBA/2J, a multipurpose neurological disease model. The Jackson Laboratory. Retrieved May 14, 2013 from http://jaxmice.jax.org/jaxnotes/512/512l.html
[8] B10.D1-Myo5a<d>/Hir. RikenBase. Retrieved May 14, 2013 from https://database.riken.jp/sw/en/B10.D1-Myo5a_d__Hir/cria315s1ria315u100000004861i/
[9] MGI - Mouse Genome Informatics. Retrieved May 15, 2013 from http://www.informatics.jax.org/
[2] New Foal for 2011-2012 Season. Delight Arab Horse Stud. Retrieved May 14, 2013 from http://delightarabhorsestud.homestead.com/Foals_.html
[3] Lavender Foal Syndrome (LFS). Animal Genetics UK. Retrieved May 14, 2013 from http://animalgenetics.eu/Equine/equine-genetic-disease/lfs.html
[4] Dereeper A., Audic S., Claverie J.M., Blanc G. BLAST-EXPLORER helps you building datasets for phylogenetic analysis. BMC Evol Biol. 2010 Jan 12;10:8. (PubMed)
[5] Dereeper A., Guignon V., Blanc G., Audic S., Buffet S., Chevenet F., Dufayard J.-F., Guindon S., Lefort V., Lescot M., Claverie J.-M., Gascuel O. Phylogeny.fr: robust phylogenetic analysis for the non-specialist Nucleic Acids Research. 2008 Jul 1; 36 (Web Server Issue):W465-9. Epub 2008 Apr 19. (PubMed)
[6] Luo, N. et al. (2012) Evidence for a role of inositol polyphosphate 5-phosphatase INPP5E in cilia formation in zebrafish. Vision Research. 75:98-107.
[7] DBA/2J, a multipurpose neurological disease model. The Jackson Laboratory. Retrieved May 14, 2013 from http://jaxmice.jax.org/jaxnotes/512/512l.html
[8] B10.D1-Myo5a<d>/Hir. RikenBase. Retrieved May 14, 2013 from https://database.riken.jp/sw/en/B10.D1-Myo5a_d__Hir/cria315s1ria315u100000004861i/
[9] MGI - Mouse Genome Informatics. Retrieved May 15, 2013 from http://www.informatics.jax.org/